ご注文方法はこちら
The LightSwitch™ Luciferase Assay System includes over 30,000 regulatory elements from the human genome that have been cloned into LightSwitch reporter vectors. With pre-cloned human promoters, lncRNA promoters and 3´UTRs available as transfection-ready LightSwitch luciferase reporter constructs, you can perform your reporter assay experiments immediately. Especially convenient are the Validated Pathway Collections, which are promoter constructs that showed significant activation or repression in response to in-house, pathway-specific induction testing. The LightSwitch System makes it easy for you to measure the transcriptional and post-transcription regulation of any gene in the genome.
To ensure high-quality results, LightSwitch reporter vectors utilize a novel, engineered luciferase gene (RenSP) and optimized reagents that provide industry-leading sensitivity. For more information, please see the tabs below as well as the Overview and RenSP Luciferase sections on the LightSwitch Luciferase Assay System page.
Find and order LightSwitch™ GoClones®
To find LightSwitch GoClone® constructs that contain the promoters of your genes of interest, use the box below to search the LightSwitch Promoter Reporter Collection. Place your order using the specific Product ID(s) you find for the clone(s) you wish to order. As we cannot stock every GoClone® in all of our offices, delivery may take 10-14 days. Click here for Search Tips »
IMPORTANT: Because all LightSwitch reporter constructs contain the optimized RenSP luciferase gene, you MUST use our LightSwitch Luciferase Assay Reagents to obtain optimal results. (Other luciferase assay reagents are not compatible with RenSP.) We also recommend that you include appropriate positive & negative control vectors when you perform your assays.
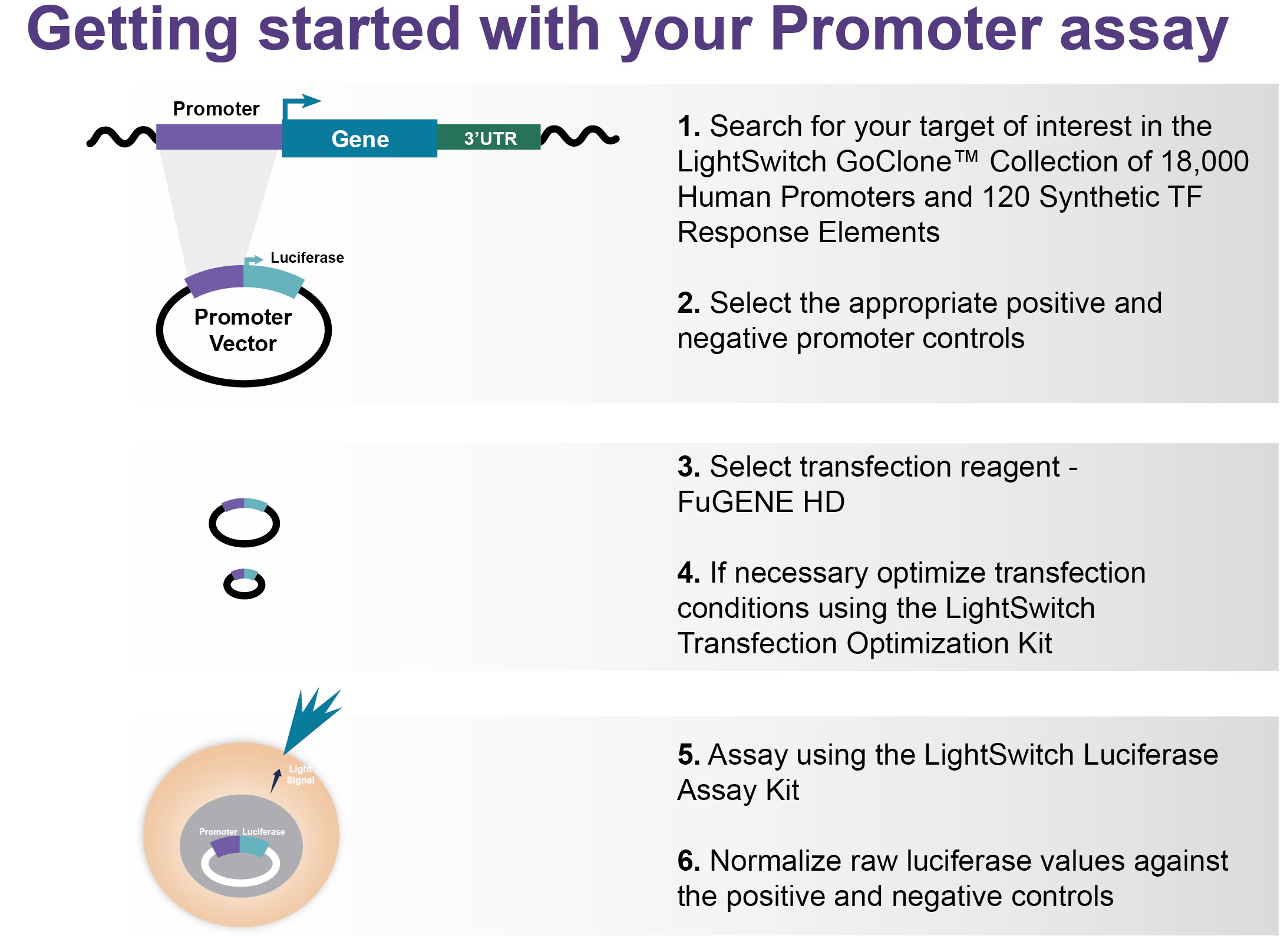
The LightSwitch Promoter Reporter Collection makes it possible to measure the functional response of promoters to stimulus in nearly any cell-based system. Plus, advances in gene assay reagent and transfection technology eliminate the need for co-transfection in most studies.
Measure promoter activity to:
- Understand the mechanism by which a gene is induced or repressed
- Characterize the functional consequences of transcription factor binding
- Measure the effect of sequence variants on promoter function

Figure 1: LightSwitch assay to measure stimulated transcription factors binding to the test promoter.
Studying transcription factor activity with the LightSwitch System
Transcription factors are important regulators of eukaryotic transcriptional regulation, and they interact with DNA-encoded regulatory elements like promoters to modulate when, where and how much of a gene’s mRNA gets made. The activity of transcription factors can be affected by the cell’s environmental conditions including exposure to signaling molecules like hormones and growth factors. Reporter assays are a powerful technique for measuring the activity of promoters in living cells. LightSwitch Reporter Constructs are created by isolating a promoter from the human genome and cloning it upstream of the RenSP luciferase reporter gene on a plasmid. Once this plasmid is transfected into a living cell, a change in promoter activity causes a change in the reporter signal (light output).

Figure 2: Response of LightSwitch SYT8 Promoter Reporter vector to various estrogen compounds.
Dose response data is shown for the SYT8 promoter (Product ID S714388) for five different estrogenic compounds. The graph summarizes the luciferase activity of this promoter in HT1080 human fibrosarcoma cells that were co-transfected with an ER cDNA expression plasmid and then treated with various concentrations of estrogen compounds for 24 hours before the luminescence was measured using the LightSwitch Luciferase Assay Kit. The Y-axis shows the relative luminescent values while the X-axis shows the molar concentration of compound on a log10 scale. The Coefficients of Variance (CVs) ranged from 10% to 12%.
The LightSwitch Luciferase Assay System and Promoter Reporter Collection are important tools for determining how a transcription factor regulates its target genes. Researchers can monitor changes in any promoter’s activity in response to variations in transcription factor function that may be modulated by techniques like over-expression (often by cDNA co-transfection; Figure 2), knock-down (often by siRNA co-transfection) or treatment with a known agonist or antagonist. LightSwitch Promoter Reporter constructs are also useful for assessing the functional consequences of transcription factor binding detected through a ChIP-based experiment and for measuring the effects of mutagenizing a putative transcription factor binding motif.
Applications
- Monitoring changes in promoter activity in response to variations in transcription factor function.
- Assessing the functional consequences of transcription factor binding detected through a ChIP-based experiment.
- Measuring the effects of mutagenizing a putative transcription factor binding motif.
- Verifying computational predictions and supplementing microarray or Next-Generation sequencing data.
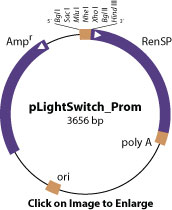
Our scientists created the LightSwitch Promoter Reporter Collection by using an algorithm to evaluate over 5 million human cDNA sequences in order to identify transcription start sites (TSS) throughout the human genome. Based on these TSS predictions, over 18,000 human promoter sequences of ~1 kb (-900…+100 bp relative to the TSS) were cloned into the pLightSwitch_Prom reporter vector, which utilizes the novel, engineered RenSP luciferase gene. After transfection, luciferase activity is assayed using the optimized regents found in LightSwitch Luciferase Assay Kit.
In addition to the over 18K endogenous promoter clones that make up the LightSwitch Promoter Reporter Collection, Active Motif also offers a collection of LightSwitch Synthetic Response Elements. These constructs contain repeats of a transcription factor binding site motif, cloned upstream of a minimal HSV-TK promoter and the RenSP luciferase gene in the Long-range Enhancer Reporter Vector, pLightSwitch_LR vector. This makes them ideal for use as a primary screen to identify compounds or conditions that affect the activity of a biological pathway. Afterward, a secondary screen can be performed using related endogenous promoter reporter constructs to detect subtle differences between compounds or conditions.
The following publications cite the use of and/or provide additional information about the LightSwitch Luciferase Assay System:
Promoters
- An antibody to amphiregulin, an abundant growth factor in patients’ fluids, inhibits ovarian tumors Carvalho et al. (2015) Oncogene
- Aspirin blocks growth of breast tumor cells and tumor-initiating cells and induces reprogramming factors of mesenchymal to epithelial transition Maity et al. (2015) Laboratory Investigation
- YB-1 regulates stress granule formation and tumor progression by translationally activating G3BP1 Somasekharan et al. (2015) Journal of Cell Biology
- Carrageenan Inhibits Insulin Signaling through GRB10-mediated Decrease in Tyr(P)-IRS1 and through Inflammation-induced Increase in Ser(P)307-IRS1 Bhattacharyya et al. (2015) Journal of Biological Chemistry
- Noncanonical STAT3 Activation Regulates Excess TGF-β1 and Collagen I Expression in Muscle of Stricturing Crohn’s Disease Li et al. (2015) Journal of Immunology
- Cytoplasmic p27 promotes epithelial-mesenchymal transition and tumor metastasis via STAT3-mediated Twist1 upregulation Zhao et al. (2015) Stem Cells
- Reduction of circulating PCSK9 and LDL-C levels by liver-specific knockdown of HNF1α in normolipidemic mice Shende et al. (2015) Journal of Lipid Research
- Functional proteomics identifies miRNAs to target a p27/Myc/phospho-Rb signature in breast and ovarian cancer Seviour et al. (2015) Oncogene
- The Endothelial Transcription Factor ERG Promotes Vascular Stability and Growth through Wnt/β-Catenin Signaling Birdsey et al. (2015) Developmental Cell
- LARP1 post-transcriptionally regulates mTOR and contributes to cancer progression Mura et al. (2014) Oncogene
- Miz-1 regulates translation of Trp53 via ribosomal protein L22 in cells undergoing V(D)J recombination Rashkovan et al. (2014) PNAS
- Arylsulfatase B regulates versican expression by galectin-3 and AP-1 mediated transcriptional effects Bhattacharyya et al. (2014) Oncogene
- Regulatory Variation in HIV-1 Dependency Factor ZNRD1 Associates with Host Resistance to HIV-1 Acquisition An et al. (2014) The Journal of Infectious Diseases
- Hypoxia-mediated downregulation of miRNA biogenesis promotes tumour progression Rupaimoole et al. (2014) Nature Communications
- NFIX Regulates Proliferation and Migration Within the Murine SVZ Neurogenic Niche Heng et al. (2014) Cerebral Cortex
- TGFβ1 Protects Cells from γ-IR by Enhancing the Activity of the NHEJ Repair Pathway Kim et al. (2014) Molecular Cancer Research
- Gli1 Protein Regulates the S-phase Checkpoint in Tumor Cells via Bid Protein, and Its Inhibition Sensitizes to DNA Topoisomerase 1 Inhibitors Tripathi et al. (2014) Journal of Biological Chemistry
- Modulation of GABAA Receptor Signaling Increases Neurogenesis and Suppresses Anxiety through NFATc4 Quadrato et al. (2014) Journal of Neuroscience
- The zinc finger transcription factor ZFX is required for maintaining the tumorigenic potential of glioblastoma stem cells Fang et al. (2014) Stem Cells
- Evidence for a novel antioxidant function and isoform-specific regulation of the human p66Shc gene Miyazawa et al. (2014) Molecular Biology of the Cell
- Functional Significance of MMP3 and TIMP2 Polymorphisms in Cleft Lip/Palate Letra et al. (2014) Journal of Dental Research
- Increased Expression of Colonic Wnt9A through Sp1-mediated Transcriptional Effects involving Arylsulfatase B, Chondroitin 4-Sulfate, and Galectin-3 Bhattacharyya et al. (2014) Journal of Biological Chemistry
- 2´-OMe-phosphorodithioate-modified siRNAs show increased loading into the RISC complex and enhanced anti-tumour activity Wu et al. (2014) Nature Communications
- Netrin-1 promotes adipose tissue macrophage retention and insulin resistance in obesity Ramkhelawon et al. (2014) Nature Medicine
- NFIB-Mediated Repression of the Epigenetic Factor Ezh2 Regulates Cortical Development Piper et al. (2014) Journal of Neuroscience
- Paeoniflorin eliminates a mutant AR via NF-YA-dependent proteolysis in spinal and bulbar muscular atrophy Tohnai et al. (2014) Human Molecular Genetics
- Cyclin-dependent kinase inhibitor 2B regulates efferocytosis and atherosclerosis Kojima et al. (2014) Journal of Clinical Investigation
- Epstein-Barr virus utilizes Ikaros in regulating its latent-lytic switch in B cells Iempridee et al. (2014) Journal of Virology
- The transcription factor Etv5 controls TH17 cell development and allergic airway inflammation Pham et al. (2014) Journal of Allergy and Clinical Immunology
- The Nutrient-Responsive Transcription Factor TFE3 Promotes Autophagy, Lysosomal Biogenesis, and Clearance of Cellular Debris Martina et al. (2014) Science Signaling
- “PTEN regulates glioblastoma oncogenesis through chromatin-associated complexes of DAXX and histone H3.3.” by Benitez et al. (2017) Nature Communications 8(15223).