<< Back to Targets & Applications Home Page
5-Hydroxymethyl Cytosine DNA Methylation
DNA methylation is an important regulator of gene expression and genomic organization. CpG methylation of gene promoters is associated with transcriptional silencing. Alterations in normal DNA methylation patterns, either local increases that result in silencing of specific cell cycle regulatory genes, or a global reduction in DNA methylation, are both involved in many different types of cancer.
CpG methylation involves the transfer of a methyl group to position 5 on the cytosine (5-methylcytosine, or 5-mC) base in a CpG dinucleotide by the action of DNA methyltransferase enzymes (DNMTs). A novel form of DNA methylation, 5-hydroxymethylcytosine (5-hmC) methylation, results from the enzymatic conversion of 5-mC into 5-hmC by the TET family of cytidine oxygenases (Figure 1). This form of methylation is prevalent in neurons and embryonic stem cells (Figure 2). While the role of 5-hmC methylation is unknown, it may represent a pathway by which 5-mC methylated DNA is converted to an unmethylated state.
Recently, two more forms of DNA methylation, related to 5-hmC methylation were discovered, 5-formylcytosine (5-fC) and 5-carboxylcytosine (5-caC). These result from iterative activity of TET enzymes on 5-hmC, further oxidizing it to 5-fC and 5‑caC. Increased levels of 5-formylcytosine and 5-carboxylcytosine are detected in the mouse male pronucleus following fertilization, which is gradually diluted by DNA replication.
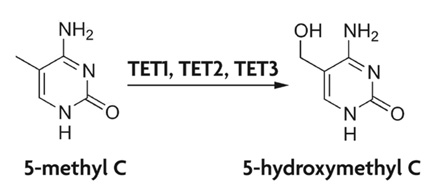
Figure 1: Schematic of conversion of 5-mC to 5-hmC.
5-methylcytosine is converted to 5-hydroxymethylcytosine by the TET family of cytidine oxygenase enzymes.
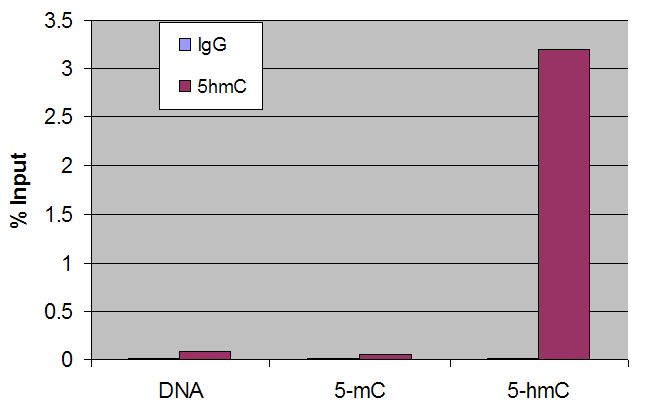
Figure 2: Dot blot using 5-mC and 5-hmC antibodies.
DNA samples were spotted onto positively charged nylon membrane and blotted with antibodies as indicated. Panel A: 5-Hydroxymethylcytidine antibody recognizing 5-hydroxymethylcytosine (1:10,000 dilution). Panel B: 5-Methylcytidine antibody (1:1,000 dilution).
Lane 1: DNA derived from mouse embryonic stem cells (150 ng).
Lane 2: DNA derived from mouse spleen (600 ng).
Lane 3: 27-base oligonucleotide containing 5-hydroxymethylcytosine (1.2 ng).
Lane 4: 33-base oligonucleotide containing 5-methylcytosine (2000 ng).
For information on products specific for 5-hydroxymethylcytosine DNA methylation, visit Active Motif's DNA Methylation & Enrichment page.
For information on 5-hydroxymethylcytosine DNA methylation services, including hMeDIP-chip and hMeDIP-Seq, visit our Custom Services page.
Download our flyer to learn more about Active Motif’s products and services to study DNA methylation.

