Anchor-Based Bisulfite Sequencing (ABBS), Enrichment-Based Nucleotide-Resolution Approach to Measure DNA Methylation
Widely-used methods to measure DNA methylation include Methylated DNA immunoprecipitation followed by sequencing (MeDIP-seq) and Whole Genome Bisulfite Sequencing (WGBS). MeDIP-seq is relatively affordable but provides semi-quantitative detection of 5-mC, whereas WGBS offers quantitative measurement of 5-mC and single-base resolution but is costly to perform.
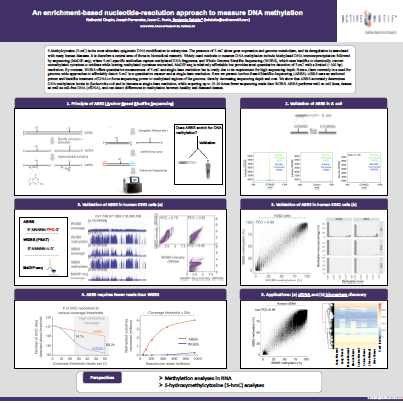
Here we present Anchor-Based Bisulfite Sequencing (ABBS), a new method using an anchored primer and bisulfite treatment of DNA to focus sequencing power to methylated regions of the genome, decreasing sequencing depth and cost. ABBS accurately determines DNA methylation levels at single-base resolution, while requiring up to 15-20 times fewer sequencing reads than WGBS, and is amenable to cell lines, tissues, and cell-free DNA (cfDNA), and can detect differences in methylation between healthy and diseased tissues.
Please note: The submission form below may not display on all browsers. For best results, we recommend using Chrome or Safari as your web browser. If you are having problems accessing the form, please contact technical support.

