<< Back to MOTIFvations Blog Home Page
Oncohistones, Gene Regulation by CpG Islands, and Epigenetics of Memory

April 2, 2019
The epigenetics field is moving quickly and hundreds of new papers are published every month, making it hard to keep up with the newest findings. We’re trying to make it a little easier for you to stay up-to-date in this exciting area by scouring the literature and bringing you short and easily digestible summaries of the most interesting and most impactful discoveries in epigenetics research each month.
This month we cover what “oncohistones” are and the role histone mutations play in cancer, new research that decouples the contributions of CpG density and DNA methylation status on the regulation of gene expression, and the epigenetic mechanisms involved in the stable maintenance of “memories” in yeast.
Epigenetics & Cancer: Histone Mutations Are Seen in Many Tumor Types
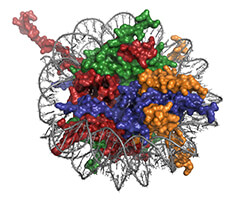
It is well known that many different cancers have mutations in genes involved in epigenetic pathways, which is why epigenetic enzymes are hot drug discovery targets. However, it’s not only mutations in epigenetic enzymes that are associated with cancer, genes encoding histones are also frequently mutated in tumors. The nature and frequency of histone mutations in cancer cells were previously poorly understood, leading Benjamin Nacev, a research fellow in David Allis’s lab, and colleagues to take a detailed look at “oncohistones.”
The team of scientists published their findings in a recent issue of the journal Nature. Previous oncohistone studies have mainly focused on a few residues on the N-terminal tail of histone H3, and this new report extended those initial papers by showing that oncogenic histone mutations are present in all four core histones and that they can be found in both the histone tails and the globular histone domains.
Furthermore, this paper presented a dataset of histone mutations functioning as common oncogenic drivers. They showed that somatic histone mutations occur in at least 4% (and probably more) of cancers from diverse tumor types.
The authors concluded in the paper that their study “should serve as a resource and starting point for the chromatin and cancer biology fields in exploring an expanding role of histone mutations in cancer.”
Reference: Nacev, B.A. et al. The expanding landscape of 'oncohistone' mutations in human cancers. Nature 567: 473-478. (2019)
Link
There’s More to DNA Methylation Sites Than the Actual Methylation
CpG dinucleotide sequences (which are sequences that contain a cytosine residue in DNA followed by a guanine) are the primary sites for DNA methylation in the genomes of mammals and many other organisms. Cytosines in most CpG sites in mammals are methylated, becoming 5-methylcytosine (5-mC), but there are also CpG dinucleotides clustered together in regulatory regions that remain unmethylated, which are referred to as CpG islands (CGIs).
These CGIs are often present in promoter regions of genes and they play important roles in regulating gene expression, likely by influencing the recruitment of RNA polymerase II and other transcription factors. While the presence of CGIs and DNA methylation modifications are both important for regulating gene transcription, it was previously not clear what the individual contributions were of the CpG dinucleotides themselves. In other words, is the sequence important, or is it the 5-mC modification that makes a difference (or a combination of both)?
In a recent paper published in Genome Research, Dominik Hartl and colleagues in the Schübeler laboratory at the Friedrich Miescher Institute for Biomedical Research in Switzerland investigated the role that CpGs played in gene regulation on a genomic scale. The researchers combined genome-wide transcription factor profiling data with high-throughput mutagenesis and reporter assays to investigate the contributions of CpG density in CGIs and DNA methylation status on the regulation of gene expression.
They found that increased CpG density correlated with increased gene expression when present in or near transcription factor binding motifs. The group of scientists also observed that overall CpG density did not impact gene expression levels, the effect was only seen in the context of certain DNA motifs.
One of the most interesting and surprising findings of the study was from experiments with DNA methyltransferase knockout cells showing that the CGI-mediated effects were independent of DNA methylation states, separating the CpG sequence contributions from the effects mediated by DNA methylation.
This study really highlights how much we still have to learn about even the most basic concepts in the epigenetic regulation of gene expression.
Reference: Hartl, D. et al. CG dinucleotides enhance promoter activity independent of DNA methylation. Genome Res. (2019)
Link
Epigenetics & Memory: How siRNA and Epimutations Help Yeast Cells Remember Their Past
They say that memory is the first thing to go as we get older. Memory research, and neuroscience in general, is one of the most fascinating areas of research. How do the different experiences we face every day shape our memories, and how does our brain know which memories to hold on to and which to let slip away?
It is difficult to study memory at a molecular level in humans and other mammals, so scientists often turn to model organisms like yeast to investigate the basic mechanisms involved behind making and remembering memories.
Lea Duempelmann and colleagues in Switzerland and the Netherlands recently described a phenomenon of transgenerational memory at the molecular level in the fission yeast, Schizosaccharomyces pombe. They were able to show that RNA-induced silencing of gene expression was mediated by epigenetic methods (trimethylation of histone H3 on lysine 9, H3K9me3) and that their yeast cells maintained a “memory” of this epigenetic silencing.
The team of researchers observed that the RNA-induced silent state of the gene encoding polymerase-associated factor 1 (Paf1) in S. pombe was stably maintained for at least 18 generations. When Paf1 was reintroduced to the yeast cells, the original silencing was lost. However, when Paf1 activity was impaired again later, the silencing was restored, even if the RNA-mediated silencing mechanism was no longer present.
While this study focused on a specific type of epigenetic memory in yeast, it's exciting to see scientists starting to unravel how organisms can remember their past experiences at a molecular level and we're looking forward to seeing what will come next.
Reference: Duempelmann, L. et al. Inheritance of a Phenotypically Neutral Epimutation Evokes Gene Silencing in Later Generations. Mol. Cell (2019)
Link
Want more epigenetics news? Check out our Epigenetics News Archive.
<< Back to MOTIFvations Blog Home Page





