Super-Enhancer Profiling Service Overview
Super-enhancers are a class of regulatory elements that participate in regulating the expression of key genes that function as master regulators of cellular identity. Not surprisingly, misregulation of these critical factors could be an important underlying mechanism in the development of diseases such as cancer. In fact, recent work has focused on the development of drugs that target super-enhancers. Super-enhancers near oncogenes are particularly sensitive to treatment and therefore show great promise as therapeutic targets.1,2,3
Let our team of ChIP-Seq experts help run the experiments and do the bioinformatics for you to help you identify super-enhancer profiles in your samples so you can publish faster and in higher impact journals.
Active Motif’s Super-Enhancer Profiling Service includes:
Customers submit frozen tissues or cell pellets, then we:
- Prepare chromatin samples and sonicate.
- Perform ChIP with a ChIP-qualified antibody.
- Construct ChIP-Seq libraries.
- Perform next-generation sequencing using the Illumina platform.
- Generate a comprehensive bioinformatics data analysis package.
- Deliver the results and publication-ready figures.
Well-Characterized, ChIP-Validated Antibodies for Known Super-Enhancer Targets:
Super-Enhancer Profiling Service Data
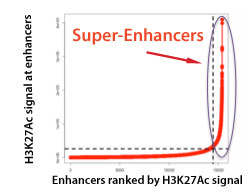
Figure 1. Identification of Super-Enhancers using Bioinformatic Analysis of ChIP-Seq Data
Enhancers are plotted in increasing order based on ChIP-Seq peak intensity. Super-enhancers are the group of enhancers above the inflection point of the curve.
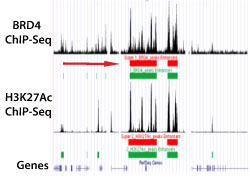
Figure 2. Super-Enhancer Profiling Using BRD4 and H3K27Ac ChIP-Seq Data
Super-enhancers in the region of the genome shown are identified by the presence of high-intensity BRD4 and H3K27Ac ChIP-Seq peaks. The super-enhancer is defined by the clustering of high-density peaks (indicated by red bars).
Super-Enhancer References:
- Hnisz et al. (2013) Cell 155, 934
- Chapuy et al. (2013) Cancer Cell 24, 777
- Loven et al. (2013) Cell 153, 320
Super-Enhancer Profiling Service Documents
Super-Enhancer Profiling Service Sample Submission Portal
Our online sample submission portal allows you to easily upload your service project samples and track your project status. Follow the sample submission instructions in the portal to ensure that all your samples arrive at Active Motif in the best possible condition and properly associated with your project.

