<< Back to MOTIFvations Blog Home Page
LINE-1 Elements: Walk the LINE-1
By Michelle Tetreault Carlson, Ph.D.
September 13, 2022
Table of Contents:
Introduction to Transposons
First discovered by Barbara McClintock in corn in the late 1940s, transposons (transposable elements), or “jumping genes” are DNA sequences that can move from one location to another in the genome. The idea was met with much suspicion at first; and understandably. What good can come from rogue fragments of DNA moving about the genome? It took decades for biologists to fully understand that not only do transposons exist in corn, but they occur in pretty much all organisms, including humans. It’s believed that transposons and their remnants speckle our genomes because they once played a critical role in species formation and evolution, by serving as a source of evolutionary novelty. Jumping fragments provided an advantage which helped organisms survive better. The legacy is that a large fraction of our human DNA (close to 50%) contains many copies of these fragments. Most of them are no longer functional or are held at bay by molecular controls. But some remain intact and become active during cell development and embryogenesis, or in response to environmental stress and aging, with the potential to wreak havoc in our genomes.
Transposons come in different lengths, ranging from just a few hundred to thousands of base pairs. They are generally categorized based on how they move about. If reverse transcription is used to copy and paste the DNA sequence from one point to another they are classified as retrotransposons. If instead transposase is used to cut and paste the DNA sequence from one place to another, they are known as DNA transposons. Both retrotransposons and DNA transposons can be further classified as either autonomous or non-autonomous, based on whether they contain the required coding sequences to make the proteins needed to move or whether they require other transposons to be mobile.
LINE-1: The Details
Long Interspersed Nuclear Elements (LINEs) are a family of autonomous retrotransposons, LINE-1, LINE-2, and LINE-3 make up the largest fraction of the human genome. Of these, the LINE-1 (or L1) elements make up the largest portion by far, accounting for about 17% of the human genome alone and with a copy number estimated at roughly half a million. Of these, most are defective, due to mutations or deletions making them incapable of retrotransposition. However, about a hundred or so of them remain intact, sometimes referred to as human-specific Line 1 (L1Hs), retaining the potential to copy and paste into new locations.
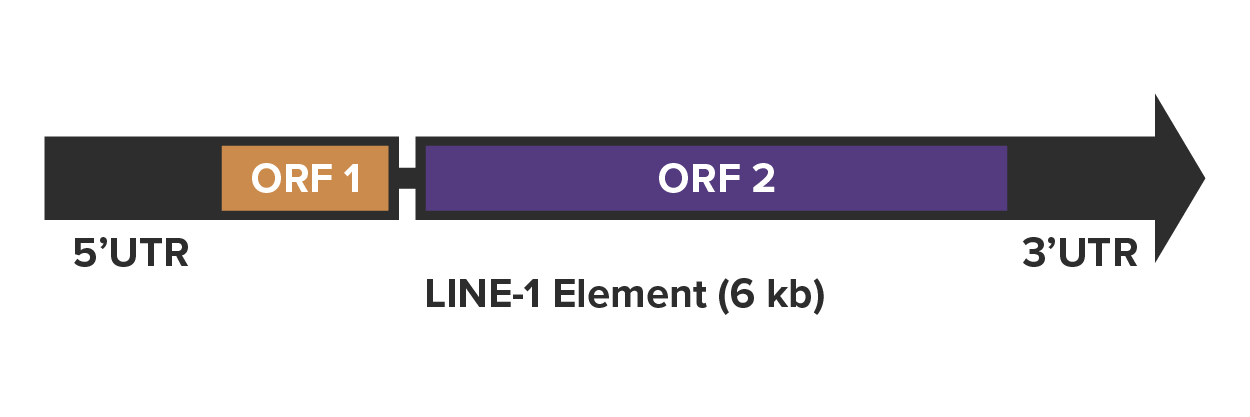
As autonomous retrotransposons, LINE-1 elements contain the coding sequences required to move themselves from one place to another and they do it via reverse transcription. The full-length LINE-1 sequence is 6000–7000 kb and contains two open reading frames, ORF1 and ORF2, which are both required for retrotransposition. These are flanked by a 5’ UTR sequence and a 3’ UTR sequence. The process of copying the LINE-1 sequence from one place to another starts with RNA polymerase II binding to the 5’-UTR promoter region of the LINE-1 sequence and facilitating the transcription of LINE-1 mRNA.
The LINE-1 mRNA is then exported to the cytoplasm where ORF1 and ORF2 are translated to ORF1p and ORF2p proteins and combined with the LINE-1 mRNA to form a ribonucleoprotein (RNP). RNP returns to the nucleus, where it interacts with DNA through a process called Target Primed Reverse Transcription (TPRT). It nicks DNA, creating a 3’ hydroxyl end that serves as a primer for reverse transcription of the LINE-1 mRNA to produce complementary DNA to the LINE-1 gene.
The LINE-1 RNP can also complex with other RNAs, inserting their complementary DNA into the genome. For example, Alu sequences, which are short interspersed nuclear elements (SINEs) of just ~300 bp long, do not contain any protein coding genes themselves. But they contain transcriptional sites for RNA polymerase III which produces Alu RNA. The LINE-1 RNP machinery can complex with Alu RNA, instigate reverse transcription to create Alu cDNA, and insert it elsewhere into the genome.
LINE-1 Repression
The hundred or so remaining LINE-1 elements that are still intact in human DNA are normally kept in check via repressive mechanisms that prevent LINE-1 gene transcription. They are kept at bay by DNA methylation, in cooperation with histone modifications, which maintain LINE-1 DNA in a heterochromatic state (the condensed, gene repressing state of chromatin). Control of jumping transposons like LINE-1 is believed to be one of the most ancient functions of DNA methylation in organisms. DNA methylation in CpG dense gene promoters has long been associated with gene silencing. Methylated cytosines are targets of methyl-CpG binding proteins which recruit repressive epigenetic factors, and they also predispose the DNA to wrap more tightly around histones.
Studies show that the full-length LINE-1 contains a ~900 bp CpG rich promoter region in the 5’UTR, with the first 100 bp most critical strongly driving mRNA transcription via recruitment of RNA polymerase II. In normal healthy somatic cells, most CpG and non-CpG dinucleotides in the LINE-1 promoter are methylated, with loss of this methylation strongly correlated with an increase in LINE-1 mRNA expression.
A number of regulatory motifs are found within the promoter region which recruit DNA-binding proteins. The LINE-1 promoter contains an initiator element (Inr) that directs transcription at the first nucleotide. Several transcription factors have been shown to be involved in both LINE-1 activation and silencing. YY1 and RUNX3 have been associated with LINE-1 activation, while E2F/Rb and MeCP2 are believed to direct LINE-1 silencing. Several other proteins including histone deacetylases (HDACs), DNA methyltransferases (DNMTs), and methyl-binding domain (MBDs) proteins act as repressors or co-repressors of LINE-1 transcription. Studies show there is post-transcriptional protein regulation of LINE-1 as well. These involve protein players functioning in RNA processing, RNP complex formation, DNA synthesis, or DNA repair.
LINE-1 and Genetic Disease
The epigenetic methylation barrier to LINE-1 activation is removed during sexual reproduction when the epigenome requires a reset. DNA methylation marks must be globally removed to allow for the adoption of the specialized, hypomethylated epigenome of the primordial germ cell and the preimplantation embryo. The result is the activation of retrotransposons like LINE-1. It has been proposed that this leads to genomic instability during these stages, as there is evidence of germline LINE-1 insertions causing sporadic cases of de novo genetic diseases. Cases of hemophilia , neurofibromatosis, and many other genetic disorders have been shown to occur as a result of LINE-1 mediated insertions of LINE-1, Alu, and other transposable element into the genome during this process.
LINE-1 and Cancer
Hypomethylation of LINE-1 promoters has long been associated with cancer. Normal somatic tissues do not express ORF1p and ORF2p, and their wayward expression contributes to carcinogenesis. As early as 1993, Thayer et. al. showed that hypomethylation of the 5’ end of LINE-1 elements in cancer cells correlates with the expression of LINE-1 proteins. LINE-1 hypomethylation occurs in the majority of cancers including, bladder and colon cancer, chronic lymphocytic leukemia, gastrointestinal stromal tumors, and lung, ovarian, and hepatocellular carcinomas.
Hypomethylation of LINE-1 has also been associated with a poor prognosis in cancer patients, especially gastrointestinal cancers. Tumor aggressiveness in 643 colon cancer samples was shown to correlate with LINE-1 hypomethylation. Poor survival in gastric cancer was linked to LINE-1 hypomethylation in more than 200 cases of gastric cancer. This trend was also found in lung, liver, esophageal, prostate, and endometrial cancers.
Activation of LINE-1 elements can drive cancer via several different mechanisms. Insertions of LINE-1 elements can disrupt the expression of tumor suppressor genes. In colon cancer LINE-1 insertions can disrupt the tumor suppressor gene APC, leading to its inactivation. In lung squamous cell carcinoma, it’s been found that a LINE-1 insertion into the tumor suppressor gene FGGY promotes tumorigenesis.
LINE-1 can also drive the expression of oncogenes. In breast cancer, Morse and colleagues first proposed that LINE-1 activation drives tumor-specific rearrangement and amplification of the oncogene MYC. In esophageal tumors LINE-1 was found to induce the amplification of CCND1 oncogene by inducing the breakage–fusion–bridge cycles. In another example, it was found that tumorigenesis in a high-grade glioma was driven by an active LINE-1 promoter that was inserted into intron 1 of oncogene FOXR2, driving overexpression.
LINE-1 and Aging
Increasing evidence suggests that repetitive elements such as LINE-1 are involved in aging and in aging related diseases. Aging results in a general loss of heterochromatin and heterochromatin-establishing factors. Consequently, the repressive mechanisms that keep LINE-1 in check, become reversed and activate LINE-1 retrotransposition. The activation of LINE-1 damages the genome directly through insertions, deletions, and recombinations. It can also spur on activation of an innate immune response, inducing autoimmunity and inflammation.
In a recent paper in Science Translational Medicine by Della Valle et. al., the correlation between LINE-1 mRNA repression, heterochromatin erosion, and the onset of aging in cells derived from patients with progeroid syndromes was examined. Progeroid syndromes make up a rare group of genetic disorders which feature symptoms of accelerated aging. They showed that LINE-1 RNA accumulation is an early event in human progeroid syndromes. LINE-1 RNA negatively regulates the enzymatic activity of the histone-lysine N-methyltransferase SUV39H1. Loss of this activity results in heterochromatin loss and the onset of senescent phenotypes in vitro. This was reversed in dermal fibroblast cells from patients with progeroid syndromes by depleting LINE-1 RNA using antisense oligonucleotides (ASOs) to the LINE-1 RNA. Depletion of LINE-1 RNA restored heterochromatin and DNA methylation, counteracting the expression of senescent phenotype cells. Antisense oligos also rescued the histophysiology of tissues and increased the life span in a mouse model of Hutchinson-Gilford progeria syndrome. The results underscore the role of LINE-1 RNA in heterochromatin homeostasis in progeroid syndromes and identify a possible therapeutic approach to treat premature aging and related syndromes.
Experiments in the Sedivy lab at Brown University demonstrated that activation of LINE-1 in senescent cells triggers a type-I interferon (IFN-I) response. IFN-I is an important contributor to an effective antiviral response, triggering the expression of interferon-stimulated genes and helping to resolve infections. However, chronic inflammation caused by IFN-I is also associated with aging (inflammaging) in senescent cells. The number of senescent cells increases as we age. Experiments demonstrated that in senescent cells the IFN-I response was specifically triggered by an increase in cytoplasmic LINE-1 cDNA. LINE-1 reverse transcription via the RNP ribonucleoprotein machinery not only produces LINE-1 cDNA in cell nuclei but can also generate LINE-1 cDNA in the cytoplasm. The immune system perceives LINE-1 cDNA in the cytoplasm as foreign DNA and instigates an IFN-I response. In older mice, which contain more senescent cells, a nucleoside reverse transcriptase inhibitor (NRTI) was used to inhibit the reverse transcriptase of LINE-1. The resulting inhibition of LINE-1 RT downregulated both IFN-1 activation and age-associated inflammation in several tissues. The authors suggest that activation of repetitive elements is an important contributor to the inflammation that is characteristic of aging. In addition, LINE-1 reverse transcription may be a target for treatment of age-related disorders.
In another study, the objective was to investigate the degree to which LINE-1 (RTP) DNA methylation in liver, kidney, and spleen tissues of mice is a marker of genetic instability and aging. In this study mice were fed either extra virgin olive oil (EVOO) or trans fatty acids (TFA) after pretreatment with the environmental carcinogen 7,12-dimethylbenz[a]anthracene (DMBA)—a harmful substance known to cause LINE-1 DNA hypomethylation. Consumption of extra virgin olive oil (EVOO) as part of a balanced diet shows preventive effects against age-related diseases and cancer, while consuming trans fatty acids (TFA) increases the risk of cardiovascular diseases. In the liver, spleen, and kidneys of these mice, both DMBA treatment and DMBA combined with TFA caused significant LINE-1 DNA hypomethylation via inhibition of DNA methyltransferase (DNMT) enzymes. However, EVOO had the opposite effect by significantly decreasing DMBA and DMBA + TFA-induced hypomethylation. Aging and LINE-1 DNA methylation are similar in human and mouse species. These results suggest that high EVOO intake may decrease the likelihood of age-related diseases and increases life expectancy because it improves the global DNA methylation pattern, reversing LINE-1 derepression.
Summary
While LINE-1 elements have been vital for fueling genetic variation and human evolution, their repression and that of other transposable elements is crucial to maintaining homeostasis in human cells. In normal healthy somatic cells, most CpG and non-CpG dinucleotides in the LINE-1 promoter are methylated, with loss of this methylation strongly correlated with an increase in LINE-1 mRNA expression. Derepression of LINE-1 is associated with many diseases, causing genomic instability, and triggering inflammation. Several different techniques for reversing the harmful effects of LINE-1 expression have been successful, including nucleoside reverse transcriptase inhibitors, antisense oligos, and dietary factors like EVOO. It’s possible that in the future, treating rogue LINE-1 expression will play a part in treating human diseases like cancer and in the reversal of aging.
References:
Kazazian HH Jr, Moran JV. Mobile DNA in Health and Disease. N Engl J Med. 2017 Jul 27;377(4):361-370. doi: 10.1056/NEJMra1510092. PMID: 28745987; PMCID: PMC5980640.
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5980640/
Zhang X, Zhang R, Yu J. New Understanding of the Relevant Role of LINE-1 Retrotransposition in Human Disease and Immune Modulation. Front Cell Dev Biol. 2020 Aug 7; 8:657. doi: 10.3389/fcell.2020.00657. PMID: 32850797; PMCID: PMC7426637.
https://pubmed.ncbi.nlm.nih.gov/32850797/
Pappalardo, X.G., Barra, V. Losing DNA methylation at repetitive elements and breaking bad. Epigenetics & Chromatin 14, 25 (2021). https://doi.org/10.1186/s13072-021-00400-z
https://pubmed.ncbi.nlm.nih.gov/34082816/
Kazazian HH Jr, Moran JV. Mobile DNA in Health and Disease. N Engl J Med. 2017 Jul 27;377(4):361-370. doi: 10.1056/NEJMra1510092. PMID: 28745987; PMCID: PMC5980640.
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5980640/
About the author

Michelle Tetreault Carlson, Ph.D.
Michelle’s interest in science was first spurred by the starry skies above her rural farm in upstate New York State, leading her to pursue a B.S. in physics. She was originally interested in astrophysics when entering the University of California, San Diego, but transitioned towards the more practical pursuit of biology earning her Ph.D. in Biophysics, studying photosynthetic proteins. Michelle’s postdoctoral research on retinal ion channels, took her further towards biology, ultimately leading to a career in the biotech industry. She enjoys chatting with scientists about their projects and interacts with them both as a Technical Support Scientist and Product Manager for Active Motif’s DNA Methylation products.
Michelle is a mother of 4 kids and 2 cats, and her hobbies include puzzles (the sign of a patient and logical mind), cooking, and pondering the human condition.
Contact Michelle with any questions at [email protected]
Related Articles
ABBS – A Novel Method to Map DNA Methylation Genome-Wide at Base Resolution
July 11, 2022
Anchor-based bisulfite sequencing (ABBS) was developed for detection of DNA methylation genome-wide. ABBS is a new technology developed by Active Motif and recently described in Communications Biology (Chapin et al., Commun Biol., 2022).
Read More
DNA Modifications Reveal the Splendor of the Plant Epigenetic Landscape
July 22, 2022
Would you like to explore the color and beauty of the epigenetic landscape of plants? Begin your journey with a discussion of recent publications highlighting the ongoing discovery and importance of DNA modifications in various plant species.
Read More
<< Back to MOTIFvations Blog Home Page








