<< Back to Targets & Applications Home Page
PARP-1: An Abundant and Ubiquitous Protein with Roles in Many Cellular Processes
by W. Lee Kraus, Cornell University
Poly(ADP-ribose) polymerase-1 (PARP-1) is an abundant and ubiquitous nuclear enzyme that plays key roles in many cellular processes, including DNA damage detection and repair, the regulation of chromatin structure, and the control of gene expression. Recent studies using a variety of experimental approaches have highlighted the molecular functions of PARP-1 in the nucleus, including gene regulation under basal, signal-activated, and stress-activated conditions2, 7, 8. PARP-1 localizes to the promoter regions of most actively transcribed genes10 and may play a role in connecting transcription to the repair of transcription-induced DNA damage6, 11. These actions of PARP-1 underlie its contributions to both normal physiology and disease processes, including inflammation, genotoxic stress, and cell death responses.
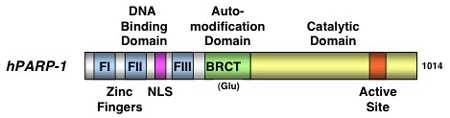
Figure 1: PARP-1 protein domains.
PARP-1 is the founding member of the PARP superfamily1. It has a carboxyl-terminal domain harboring an enzymatic activity that catalyzes poly(ADP-ribosyl)ation reactions using nicotinamide adenine dinucleotide (NAD+) molecules as a donor of ADP-ribose groups2, 7 (Figure 1). PARP-1 also has an amino-terminal DNA binding domain (DBD) containing three zinc finger motifs, a nuclear localization signal, and an automodification domain (AMD) that functions as the target of covalent auto-poly(ADP-ribosyl)ation1, 2 (Figure 1). Together, these domains allow PARP-1 to interact with genomic DNA and chromatin, poly(ADP-ribosyl)ate a variety of nuclear target proteins, and regulate nuclear functions.
Numerous nuclear targets for PARP-1 enzymatic activity have been identified, including core histones, the linker histone H1, and a variety of transcription factors. PARP-1, however, is the major acceptor for poly(ADP-ribosyl)ation reactions in vivo2, 9. PARP-1 associates with a variety of nuclear complexes containing DNA repair- and transcription-related factors4. These interactions are critical for PARP-1 function. PARP-1 is cleaved by caspases during apoptosis, and the PARP-1 cleavage product is a reliable indicator of apoptotic responses. Recent biochemical, genomic, proteomic, and cell-based studies have highlighted the role of PARP-1 in these processes and provided new insights about the molecular mechanisms governing PARP-1 actions2, 7, 8. These studies have also demonstrated how PARP-1 functions as an integral part of cellular signaling pathways that culminate in genomic outcomes8. These insights have led to the development of a variety of PARP inhibitors that have shown promise for the treatment of cancer, ischemic injury, and cardiovascular disease3, 5.
PARP-1 Interactions and Targets
PARP-1 exhibits physical and functional interactions with (1) core histones, (2) the linker histone H1, (3) topoisomerases I and II, and (4) DNA binding transcription factors such as NFκB, B-MYB, Oct-1, nuclear receptors, YY1, TBP, and the HTLV Tax-1 protein. PARP-1 also associates with a variety of regulatory complexes, including a TLE (transducin-like enhancer of split) corepressor complex, a Mediator coregulator complex, a condensin I/XRCC1 repair complex, a macroH2A1.1 nucleosome complex, and a CTCF insulator complex8. Many of these proteins are also targets for poly(ADP-ribosyl)ation by PARP-1.
References
- Amé, J.C., Spenlehauer, C., and de Murcia, G. (2004) The PARP superfamily. Bioessays 26: 882-893.
- D‘Amours, D., Desnoyers, S., D‘Silva, I., Poirier, G. (1999) Poly(ADP-ribosyl)ation reactions in the regulation of nuclear functions. Biochem. J. 342: 249-268.
- de la Lastra, C.A., Villegas, I., and Sanchez-Fidalgo, S. (2007) Poly(ADP-ribose) polymerase inhibitors: new pharmacological functions and potential clinical implications. Curr Pharm Des 13: 933-962.
- Droit, A., Hunter, J.M., Rouleau, M., Ethier, C., Picard-Cloutier, A., Bourgais, D., and Poirier, G.G. (2007) PARPs Database: A LIMS systems for protein-protein interaction data mining or Laboratory Information management system. BMC Bioinformatics 8: 483.
- Horvath, E.M., and Szabo, C. (2007) Poly(ADP-ribose) polymerase as a drug target for cardiovascular disease and cancer: an update. Drug News Perspect 20: 171-181.
- Ju, B.G., Lunyak, V.V., Perissi, V., Garcia-Bassets, I., Rose, D.W., Glass, C.K., and Rosenfeld, M.G. (2006) A topoisomerase II beta-mediated dsDNA break required for regulated transcription. Science 312: 1798-1802.
- Kim, M.Y., Zhang, T., and Kraus, W.L. (2005) Poly(ADP-ribosyl)ation by PARP-1: 'PAR-laying' NAD+ into a nuclear signal. Genes Dev 19: 1951-1967.
- Kraus, W.L. (2008) Transcriptional control by PARP-1: chromatin modulation, enhancer-binding, coregulation, and insulation. Curr Opin Cell Biol 20: 294-302.
- Kraus, W.L., Lis, J. (2003) PARP goes transcription. Cell 113: 677-683.
- Krishnakumar, R., Gamble, M.J., Frizzell, K.M., Berrocal, J.G., Kininis, M., and Kraus, W.L. (2008) Reciprocal binding of PARP-1 and histone H1 at promoters specifies transcriptional outcomes. Science 319: 819-821.
- Perillo, B., Ombra, M.N., Bertoni, A., Cuozzo, C., Sacchetti, S., Sasso, A., Chiariotti, L., Malorni, A., Abbondanza, C., and Avvedimento, E.V. (2008) DNA oxidation as triggered by H3K9me2 demethylation drives estrogen-induced gene expression. Science 319: 202-206.
PARP-1 Inhibition, DNA Repair and Epigenetics
PARP-1 is one of the most abundant proteins in mammalian cells, with a typical number of 1-2 million copies of protein per cell. PARP-1 has an N-terminal domain that binds with high affinity to double-stranded breaks, cruciforms and crossovers in DNA, and also to nucleosomes. Binding to DNA is associated with activation of the C-terminal catalytic domain, which PARylates numerous proteins, as well as PARP-1 itself. One of the first targets of PARP-1 upon binding DNA double-strand breaks is the kinase ATM, which initiates the DNA damage repair response, including phosphorylation of histone variant H2AX at Serine 139, and recruitment of BRCA1 and 2, Rad51, and the other proteins involved in DNA repair. In the new report from Fong et al., PARP-1 inhibitor Olaparib (AZD2281) was given to breast, ovarian, and prostate cancer patients in a phase I trial, resulting in a very high partial or complete response rate for those patients with genetic lesions in BRCA1 and 2, who are most susceptible to these tumors. The mode of action is thought to involve the role of PARP-1 in DNA repair, and is highly plausible given the necessity of PARP-1 to activate ATM, and previous observations that inhibition of PARP-1 killed cells deficient in the homologous recombination/Rad51-dependent DNA repair process. However, PARP-1 has effects on other proteins that merit investigation for possible contributory efficacy. From an epigentics perspective, PARP-1 has been shown to regulate DNA methylation, possibly through regulation of DNMT1, and modulation of the binding of CTCF to insulator elements.
Active Motif Products Related to PARP-1
Active Motif offers a large number of antibodies and proteins directed against PARP-1 and other proteins that interact with PARP-1. Click on the links in the table below for complete details. In addition, we offer our Chromatin Assembly Kit, which makes possible in vitro packaging of your circular or linear DNA in natural, ordered nucleosome arrays; this enables you to perform in vitro chromatin IP (ChIP), transcription and histone modification assays of your sequence of interest in a native chromatin environment.
| PARP-1-related Proteins & Antibodies | Applications |
|---|---|
| Recombinant BRCA1 protein | DNA repair, transcription, protein-protein interaction, and cell growth assays |
| Recombinant Rad51 protein | DNA-protein and protein-protein interaction assays |
| ATM phospho Ser1981 mouse monoclonal (Clone 10H11.E.12) | IF |
| Aurora B rabbit polyclonal antibody | IF, WB |
| Caspase-3 mouse monoclonal antibody | WB |
| DNMT1 mouse monoclonal antibody | ChIP, IHC, IP, WB |
| Histone H2A phospho Ser129 rabbit polyclonal antibody | ChIP, IF, IP, WB |
| Histone macroH2A1 rabbit polyclonal antibody | IF, WB |
| Histone H2A/H4 phospho Ser1 rabbit polyclonal antibody | WB |
| Histone H2AX phospho Ser139 rabbit polyclonal antibody | IF, WB |
| p53 rabbit polyclonal antibody | WB |
| PARP-1 N-terminal rabbit polyclonal antibody | WB |
| PARP-1 C-terminal rabbit polyclonal antibody | WB |
| Rad21 mouse monoclonal antibody (Clone 52A311) | WB |
| Rad51 rabbit polyclonal antibody | IF, IHC, IP, WB |
Application Key:
- ChIP = Chromatin Immunoprecipitation;
- ELISA = Enzyme-linked Immunosorbent Assay;
- EMSA = Electrophoretic Mobility Shift Assay;
- FACS = Flow Cytometry;
- IF = Immunofluorescence;
- IHC = Immunohistochemistry;
- IP = Immunoprecipitation;
- WB = Western blot

