<< Back to MOTIFvations Blog Home Page
最良のChIPデータを得るための手引き

March 15, 2019
クロマチン免疫沈降法(ChIP, chromatin immunoprecipitation)は、DNA結合タンパク質とクロマチンの相互作用や、ゲノム全体にわたるヒストン修飾の局在を解析するために最も広く用いられる実験手法の一つとなった。ChIPは多くの分子生物学者にとって日常的な手法となったが、そのプロトコルには複雑な手順が多く含まれており、実験の成功には各段階で克服すべき複数の困難が伴う。
ChIPアッセイで毎回ベストな結果を得る方法
25年以上にわたり、Active Motifは遺伝子調節とタンパク質機能の研究用キットおよびアッセイ法の開発における先駆者であり続けてきました。当社は15年以上前に世界で初めてChIPキットを開発し、以来改良と革新を重ねています。現在、Active Motifは、最高品質のChIP結果を可能な限り容易かつ迅速に生成するための様々なChIPキットや関連製品、およびChIP-Seq受託サービスを提供しています。
当社は、世界中のどのグループよりも多くのChIPおよびChIP-Seq実験を実施しており、長年にわたり豊富な知見を得てきました。皆様のChIPアッセイで最良の結果を得るための、当社が厳選した秘訣を喜んで共有いたします!
ChIPアッセイの歴史

ChIPアッセイにはすでに長い歴史があります。今日我々が知るChIPへとつながる実験的手法は、1980年代にコーネル大学のJohn Lisの研究室で初めて記述されました。初期のChIP実験はRNAポリメラーゼIIのゲノム局在を調べたもので、当初は細菌細胞で、後にショウジョウバエ細胞で実施されました。これらの初期の実験はハイブリダイゼーションに基づくアッセイにより解析された。
クロマチン免疫沈降法はその後大きく発展しました。このアッセイは、多くの研究室で様々な生物種やサンプルタイプに対して広く用いられる技術となり、多数の論文や重大な発見につながっています。ChIPアッセイの下流解析も、エンドポイントPCRから定量PCR (qPCR)に移行し、さらに近年では次世代シーケンシングへと進化を遂げています。
ChIP実験における5つの重要なステップ
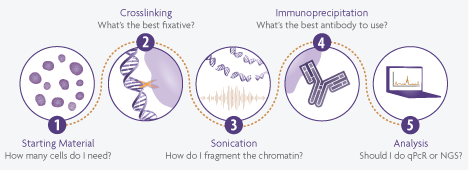
すべてのChIP実験において考慮すべき重要な観点として、以下の5つが挙げられます。
- まず、最も基本的な点として、ChIPアッセイを実施するための出発材料が必要です。細胞や組織のどちらも使用可能であり、それぞれの必要量は状況に応じて異なります。
- 次に、細胞を固定してタンパク質とDNAを架橋させるのが一般的です。ほとんどのプロトコルではホルムアルデヒドを使用しますが、固定条件や添加剤にはいくつかのバリエーションが存在します。
- 続いてクロマチンを断片化しますが、通常は超音波処理(ソニケーション)が行われます。
- その後、実際の免疫沈降(IP)ステップがあります。使用する抗体が極めて重要ですが、磁気ビーズとアガロースビーズのどちらを使用するかの判断も同様に重要です。
- 最後に下流解析を行いますが、近年は通常、qPCRまたは次世代シーケンシングのいずれかです。
これらの重要な各ステップについては、以下で詳細に説明します。また、このトピックについてはActive Motif Epigenetics Podcastでも取り上げていますので、読むよりも聴く方がお好みの方はそちらもお試しださい。
ChIPアッセイに必要な細胞数

ChIPアッセイの計画を立てる際に最初に考慮すべきことは、出発材料です。どのサンプルタイプを使用すべきか、またどの程度の細胞数(あるいは組織量)を使用すべきでしょうか?
従来、各ChIP実験には数百万細胞が使用されてきました。この大量の細胞を必要とする要件は、特に初代細胞や幹細胞など十分な細胞数を確保することが困難なサンプルを用いて良好なChIPおよびChIP-Seq結果を得る上で最大の障壁でした。
少数の細胞でChIPは実施可能か?
近年、少数の細胞に特化したChIPキットが開発されたことで、ChIPに要する細胞数は大幅に減少しました。これらのキットにより、研究者は数百万ではなく数千個の細胞で実験が可能になり、さらに細胞数を削減する新技術が日々登場しています。
ChIP用サンプルを適切に固定・調製する方法
ChIP反応では様々なサンプルタイプが使用可能ですが、すべてのChIPプロトコルにおいて重要な要素は、実際に細胞をどのように処理するかです。ほとんどのChIP実験では、DNAに結合するタンパク質を架橋するためにサンプルを固定します。そして、異なる細胞や組織では異なる固定のプロトコルが必要です。考慮すべき重要な点は、使用する固定剤の種類と、その反応を停止するまでの固定時間の長さです。
ホルムアルデヒドはChIPにおける最も標準的な固定剤
標準的な固定剤は通常ホルムアルデヒドです。一般的な実験室にあるホルムアルデヒドは通常、安定化剤としてメタノールが添加された重量比率37%のホルムアルデヒド溶液(ホルマリン)です。しかし、メタノールが含まれる問題点の一つとして、細胞膜の透過性を高める可能性が挙げられます。これにより、メタノールを含まないホルムアルデヒドを使用する場合よりも多くのホルムアルデヒドが細胞内に取り込まれ、最終的にクロマチンの過剰固定を引き起こす可能性があります。一方、メタノールなしではホルムアルデヒドは安定性が低く重合を起こすため、用時調製しないと効果が低下します。
特定の実験には他の固定剤も効果的ですが、ホルムアルデヒドが最も一般的に使用されるのは、非常に近接した2つの分子を共有結合により架橋できるためです。ただし、そのような結合を避けたい場合には、DSG (N,N‘-Disuccinimidyl Glutarate)のような他の固定剤を使用することにより、やや離れた距離にある2つの分子を架橋することが可能です。これらはすべてChIP実験をデザインする際に考慮すべき点ですが、ほとんどの場合、ホルムアルデヒドは固定条件として非常に優れており、多くの論文において採用されています。
ChIP実験において適切な細胞の固定時間
固定法の次に検討すべき点は、細胞を固定する時間の長さです。ほとんどのプロトコルでは、最終濃度1%のホルムアルデヒドを用いて10~15分間固定する条件が指定されています。これらの固定条件は多くのサンプルにおいて有効ですが、場合によっては最適化が必要なこともあります。
もし、用いる組織片が大きい場合、懸濁液中のすべての細胞が液体に面しているとは限りません。そのため、ホルムアルデヒドをそれらの細胞に浸透させるには固定時間の延長が必要になることがあります。
また、アッセイにホルムアルデヒドを含まないプロトコルも存在し、これはNative ChIPと呼ばれます。Native ChIPアッセイでは、DNAと対象タンパク質が人為的に架橋されていない状態で解析を行います。この方法は一般的に、ヒストンのようにDNAと非常に強く結合しているタンパク質に対してのみ有効です。
ソニケーションによるクロマチンの断片化と可溶化
細胞を固定した後は、免疫沈降反応用の可溶化したクロマチンを得るために細胞を溶解し、さらにソニケーションを行う必要があります。一般的には、クロマチン断片の長さが200〜1,000塩基対になるようにするのが望ましく、適切な断片を得るためにはいくつかの点に注意する必要があります。
固定後の核内クロマチンへのアクセス
細胞を固定した後、クロマチンを取り出すためには、まず核の内部へアクセスする必要があります。さらに、クロマチンに到達するだけでなく、抗体により効率よく回収するために、クロマチンを十分細かく断片化することも必要です。
必須の工程ではありませんが、一部の有力なクロマチン生物学の研究室では、クロマチンを断片化する前に、まず核を単離する方法が採られています。細胞の固定後に実施可能な簡便な核単離プロトコルもいくつか知られています。核を単離した後、それをより強い条件の溶解バッファーに再懸濁すると、核が開き、内部のクロマチンにアクセスできるようになります。
クロマチンの断片化―どの方法が最良か?
クロマチンにアクセスしたら、DNAを小さく断片化する必要があります。これにはいくつかの方法がありますが、推奨される手法はソニケーションです。核をドデシル硫酸ナトリウム (Sodium Dodecyl Sulfate)を含む溶解バッファーに再懸濁した後、細胞に超音波を照射します。この処理を行う装置は数多く存在します。本質的には、DNA断片を全て200~1000塩基対の長さに分断するという考え方です。
クロマチン断片は大きすぎても小さすぎてもいけません。言うまでもなく、細胞のソニケーションでは、固定されたクロマチン内のDNAやタンパク質が比較的強い超音波にさらすことになります。処理時間が長すぎると、抗体が認識できないほどタンパク質やDNAが分解される可能性があります。また、ソニケーションで発生する熱によりタンパク質が変性し、使用している抗体が標的とするエピトープを認識できなくなる場合もあります。
ChIPアッセイで最良な結果を得るためのソニケーション条件の最適化
ChIPプロトコルの他の工程と同様に、あらゆるサンプルに最適な単一のソニケーション条件は存在しません。最適なソニケーション条件は細胞種や組織によって異なり、使用するサンプルに最適化する必要があります。
アクティブ・モティフではChIPアッセイを実施する際、ほとんどのサンプル種に対応する基本的なソニケーション条件を用意しています。ただし、追加のソニケーションを必要とする細胞種も数多く存在します。例えば、T細胞はソニケーションが非常に困難な傾向があります。これはおそらく、血液中を浮遊し様々な物理的ストレスに耐える必要性から頑丈に作られているためでしょう。また、この細胞は非常に小さく高密度で開砕が難しいため、最良の結果を得るには特別な溶解、およびソニケーション条件が必要となります。
酵素を用いたクロマチンの断片化法
ソニケーションは非常に効率的でランダムに断片化できるため、ほとんどの研究者はDNAの断片化にソニケーションを好んで使用しますが、考慮すべき潜在的な欠点も存在します。ソニケーションの欠点の一つは、処理しすぎるとタンパク質を損傷したり、タンパク質-DNA相互作用を破壊する可能性があることです。
クロマチンをもう少し穏やかに扱いつつ適切なサイズまで断片化したい研究者は、酵素的なせん断法を用います。もっとも一般的な酵素的せん断法は、ミクロコッカスヌクレアーゼ(MNase)の使用です。MNaseはATリッチな配列を切断する酵素であり、反応時間を調節することにより目的のサイズのクロマチン断片を得ることができます。
MNaseを長時間処理すると、ヒストンやヌクレオソームのサイズ(約146塩基対)を最小単位とした非常に小さな断片が得られます。一方、処理時間をごく短くすると、ChIPで推奨される約1,000塩基対程度の断片を含め、さまざまなサイズの断片が得られます。
しかし、多くの研究者がクロマチン断片化にMNaseを用いない主な理由は、酵素による切断に偏り(バイアス)が生じやすいためです。MNaseは特定のDNA配列を他よりも優先的に切断する傾向があるため、この酵素による断片化は、解析結果にバイアスがかかることが知られています(訳注:MNaseには認識配列のような配列特異性はありませんが、AT-rich領域を切断しやすい性質があります)。
ChIPアッセイに最適な抗体の種類とは?
クロマチンを単離して、最適なサイズになるようソニケーションしたら、免疫沈降へ進みます。サンプル調製やソニケーションの工程がどれほど完璧でも、適切な抗体を使用しなければ良好なChIPの結果は得られません。ChIPアッセイで使用可能な抗体には、ポリクローナル抗体、モノクローナル抗体、リコンビナント抗体など複数のタイプがあります。ここでは、それぞれの抗体の利点を紹介し、実験に最も適した抗体を選ぶための参考にしていただきたいと思います。
ChIPに最適な抗体はどれかという点については、これまでこの分野で長く議論が続いてきました。ポリクローナル抗体の方がモノクローナル抗体より優れているのでしょうか。実際のところ、その答えは状況によって異なります。さらに近年では、リコンビナント抗体という新たな選択肢も登場し、この議論に新しい側面が加わっています。Active Motifのような抗体を製造・販売する企業では現在、抗体をリコンビナントタンパク質 として生産しており、従来のモノクローナル抗体よりも抗体集団の均一性が高い場合が多くなっています。
あらゆる条件において、特定のタイプの抗体が他よりも優れていると断言することは非常に困難です。
ChIP抗体は抗体のタイプよりも特異性が重要
ChIP用抗体を選択する際に評価すべき最も重要なことは、標的タンパク質を固定化する条件において高い特異性をもって認識できるか、そして免疫沈降(IP)と洗浄ステップを通じて結合を維持できるかどうかです。
ChIP法が開発された初期の頃は、多くの研究者がポリクローナル抗体の使用を好みました。これは、ポリクローナル抗体が複数のエピトープを認識する抗体の混合物であるため、ホルムアルデヒドによる固定の影響を受けない抗体も含まれる可能性があるからです。
しかし現在では、ほとんどの標的タンパク質に対して複数の異なる抗体が利用可能であり、ChIP向けに検証されていれば、ポリクローナル抗体、モノクローナル抗体、リコンビナント抗体のいずれもChIPアッセイで良好な性能を発揮することが明らかになっています。
標的タンパク質に対するChIP検証済み抗体がない場合の対処法
前述のように、ChIP実験を成功させる上で最も重要なステップの一つは、目的のタンパク質に対して適切な抗体を見つけることです。これは多くの研究者にとって大きな課題となる場合があります。DNA結合タンパク質やヒストン修飾のうち、よく研究されているものについては複数の抗体が市販されていますが、必ずしもすべてのターゲットでそうとは限りません。
ある抗体がウエスタンブロットや他のアッセイで良好に機能することが確認されていても、それがChIPでも同様に機能するとは限りません。ChIP実験に適した抗体を見つける最も確実な方法は、まず文献を調べることです。最初に行うべき基本的なアプローチは、同じ標的タンパク質またはヒストン修飾に対してChIP解析を行った著名な研究者の論文を探すことです。
もし、そのような抗体が見つからない場合には、ChIPと同様のクロスリンク(固定)を行う実験条件で使用された抗体の論文を探してみると良いでしょう。具体的には、免疫蛍光染色(IF, Immune Fluorescence)や免疫組織化学染色(IHC, Immunohistochemistry)です。これらの手法はいずれも固定条件下のサンプルに対して行われるため、これらのアッセイで機能する抗体は、ChIPにも利用できる可能性があります。ただし、IFやIHCでは標的タンパク質のエピトープを抗体が認識できたとしても、ChIPアッセイに特有の条件により抗体の結合能を阻害する可能性があるため注意が必要です。
例えば、ChIPでクロマチンを可溶化・断片化するために使用されるバッファーには、多くの場合、比較的強い陰イオン性界面活性剤SDSが含まれています。SDSが高濃度で存在すると抗体とタンパク質の結合が阻害されるため、抗体が理論上は抗原を認識できたとしても、免疫沈降に必要なほど強く結合することができない場合があります。このような条件により、目的のタンパク質-DNA複合体を十分に回収できず、ChIPがうまく機能しないことがあるのです。
ChIP実験で使用した抗体が機能したかどうかをどう判断するか?
ChIP反応の結果を信頼できるものにするためには、適切なコントロール実験を行うことが極めて重要です。最良のアプローチは、対象となる因子やヒストン修飾が存在することが既知のゲノム領域(ポジティブコントロール)と、存在しないことが既知の領域(ネガティブコントロール)に対して、それぞれ特異的なqPCR反応を行うことです。
ネガティブコントロールとして、標的タンパク質に対する抗体とは無関係な正常IgGを用いてChIP反応を行うことを好む研究者もいます。この考え方では、この抗体で検出されるシグナルは、対象分子に特異的ではないためバックグラウンドだとみなされます。この方法は、コントロールをまったく設定しないよりはよいものの、あまり理想的ではありません。というのも、ChIPで使用する特異的抗体は正常IgGとは異なるバックグラウンド特性を示す可能性が高く、厳密には公平な比較にはならないからです。
免疫沈降に最適なビーズとは?
免疫沈降の後、対象の抗体で標識されたタンパク質-クロマチン複合体は、protein Aまたはprotein Gを結合させたビーズと反応させて回収します。利用可能なビーズにはいくつかの種類がありますが、「ChIPに最適なビーズ」は分野内でも意見が分かれるテーマです。
多くの研究者は磁気ビーズを好んで使います。磁気ビーズにはいくつか利点があり、洗浄工程を非常に素早く簡単に進められること、そして磁石機構を備えた装置があれば自動化にも対応しやすいことが挙げられます。また、チューブの側面にビーズが磁石で引き寄せられるのを目視で確認できるため、洗浄バッファーを除去する際に免疫沈降したサンプルを誤って吸引するのを防ぐことができます。
磁気ビーズの代替としてはアガロースビーズがあります。こちらにも独自の利点があり、特にバックグラウンドが低く、感度が高いことが知られています。アガロースビーズでは各工程ごとに遠心分離が必要なため、洗浄に手間と時間がかかることが問題でした。しかしActive Motifは、使いやすいカラム方式のprotein Gアガロースビーズを採用し、この問題を解決しました。すべての洗浄をカラム上で迅速かつ簡単に行うことができます。カラムに洗浄バッファーを加えるだけで液が流れ去るため、洗浄のたびにビーズを遠心する必要はなく、多数のChIP反応を並行して行うことも可能です。
ChIPの結果を解析するための最良の方法とは?
ChIP反応の最終段階は、免疫沈降したDNAの解析です。クロスリンクを解除し、結合タンパク質の消化してDNAを精製した後、研究者は通常、PCRまたは次世代シーケンシング(NGS)を用いて、単離したDNA断片を解析します。すでに興味のある遺伝子座が分かっていて、さらに解析対象の領域数がそれほど多くない場合には、qPCRによる解析が適しています。一方、バイアスのない方法で、目的の因子やヒストン修飾がゲノム上のどこに存在するのかを網羅的に調べる場合には、NGSが有効です。
qPCRは、ChIPにおいて対象である因子が結合している領域を解析する初期の手法であり、現在でも多くの研究者に使われています。最大の制約は、解析対象となる結合領域を事前に特定し、その領域を特異的に増幅するためのプライマーを設計しなければならない点です。qPCRベースの解析は、異なる実験条件間における既知の配列に対する結合の違いを比較する場合には有効ですが、一般的に、対象とする因子の新たな結合部位を見つけることを目的とした実験には適していません。
ChIP-seq のようなNGSベースの解析では、1回のChIP実験から圧倒的な情報量を得られる反面、NGSライブラリーを作製するための追加の処理が必要なことに加え、結果を解釈するためのバイオインフォマティクス解析も必要になります。このことは多くの研究者にとって障壁となっています。NGSはqPCRベースの解析よりも割高ですが、そのコストは急速に低下しており、qPCRに比べて飛躍的なデータを得られることから、ChIP解析はNGSベースの解析への移行が加速しています。さらに、ハイインパクトな学術誌の中には、論文掲載にChIP-seqを求めるところも増えています。
比較的使いやすいオープンソースの解析プラットフォームの登場により、バイオインフォマティクスは以前より手軽になっていますが、ほとんどのChIP-Seqプロジェクトでは、専門のバイオインフォマティシャンが解析を担当し、データの解釈や可視化のための図表作成まで担うことが一般的です。
結論:ChIPアッセイの困難は克服可能!

ChIPアッセイには多くの工程があり、そのすべてが実験全体の成功にとって極めて重要です。しかし、綿密な計画、適切なChIPキットの選定、ChIP検証済み抗体, using ChIP-validated antibodiesの使用、そしていくつかの最適化を行うことにより、ChIPアッセイにおける数々の困難は克服することができます。
また、最高品質のChIP-Seqのデータをできるだけ早く必要とする研究者の方には、”end-to-end”のChIP-Seq受託サービスの利用をお勧めします。このサービスを利用すれば、細胞や組織をアクティブ・モティフ日本法人に送るだけで、数週間以内に解析済みのデータを受け取ることができます。
About the author

Kevin Bryant, Ph.D.
Kevin was born in Northern California, in the small town of Eureka, but his studies brought him to the East Coast for about a decade where he came to love Boston sports teams and ended up meeting his wife. Now back home in his native California, he likes to explore the San Diego area with his family and friends. When not working at Active Motif or shopping at Costco, Kevin enjoys trying to become a wine connoisseur and is always searching for a beer he likes better than Pliny the Elder.
Contact Kevin on LinkedIn with any questions, complaints, or compliments – or to get IPA recommendations.
Related Articles
Complete Guide to Sonication of Chromatin for ChIP Assays
January 31, 2020
Chromatin shearing/sonication is one of the most important steps in the ChIP workflow and is generally the step that exhibits the most variability. This article discusses the differences between each available sonication method and highlights their pros and cons.
Read More
Comprehensive Guide to Understanding and Using CUT&Tag Assays
March 9, 2020
CUT&Tag shows a lot of promise and has the potential to alleviate some ChIP limitations, but it also has its own set of limitations that must be considered. This article covers what CUT&Tag is and describes the advantages and drawbacks of this method.
Read More
<< Back to MOTIFvations Blog Home Page










